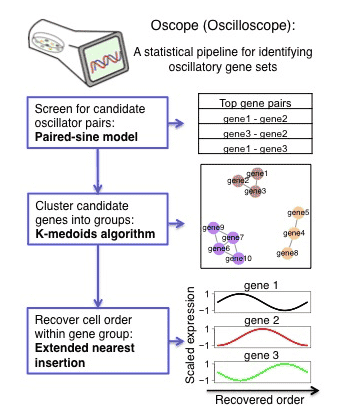
A statistical pipeline for identifying oscillatory genes in unsynchronized single cell RNA-seq experiments
Introduction
Download
Oscope was built under R version 3.2.1. R is a free and popular statistical software package.
Installation
To run Oscope, start R and within R type:
source(“http://bioconductor.org/biocLite.R”)
devel = “http://bioconductor.org/packages/3.2/bioc”
biocLite(“Oscope”, siteRepos = devel, type=”source”)
library(Oscope)
packageVersion(“Oscope”)
demo(Oscope)The demo runs through many of the commands contained in the vignette automatically, using simulated data for illustration (it will take a few minutes). To apply Oscope to a particular dataset, the commands within the vignette may be followed once that dataset is read into R.
Vignette
References
Leng, N.*, Chu, L.-F.*, Barry, C., Li, Y., Choi, J., Li, X., Jiang, P., Stewart, R. M., Thomson, J. A., and Kendzrioski, C. Oscope identifies oscillatory genes in unsynchronized single cell rna-seq experiments. Nature Methods, accepted
Updates
- Latest updates could be found at NEWS on Bioconductor
- 06/24/2015 Oscope 0.99.1 is available on Bioconductor now!
- 06/24/2015 Oscope 0.0.6 is released