Positive-strand RNA viruses comprise a vast and often fearsome roster that includes poliovirus, hepatitis C, dengue, Zika, HIV, SARS-CoV-2, and many agriculturally important plant viruses. But it’s the small things — the molecular machines fueling viral replication — that lie behind their immense global impact.
New research from Morgridge scientists reveals just how small.
From neck to crown to elbow

Positive-strand RNA viruses replicate their genomes in special compartments called replication organelles, which organize the process and shield it from host detection. Using a model nodavirus, the lab of Paul Ahlquist has revealed many aspects of how viruses induce these organelles by folding host membranes into new vesicles with narrow, neck-like connections to the surrounding cell environment. Through these necks, viruses can import material needed from the host and export newly synthesized viral RNA. Using cryo-electron tomography, they provided the first visualization of all components of such a replication organelle — not only membranes but also the viral RNAs being replicated and the viral proteins doing the replication. And they discovered that these viral proteins were organized in a 12-fold symmetric ring structure gating the neck of the replication organelle, which they named the “crown.” Recent studies with two other viruses show that crown structures built on similar principles are used by many if not all positive-strand RNA viruses, making crown structure, assembly, and operation of great interest.
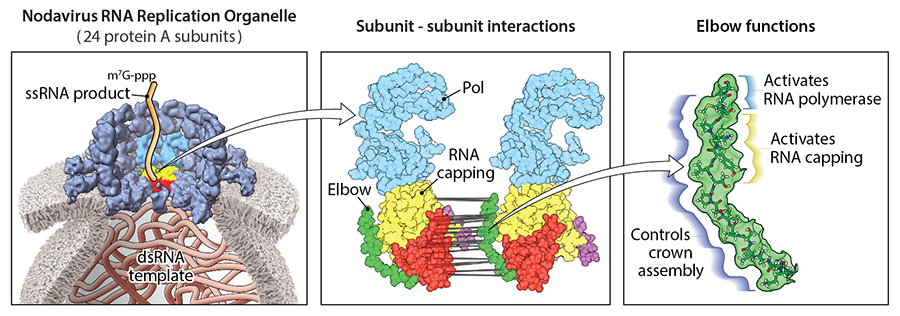
The nodavirus crown is built from two stacked rings, each bearing 12 copies of a large, multi-domain viral protein called protein A. With the Tim Grant Lab at Morgridge, the researchers produced a detailed, near atomic-resolution cryo-electron microscopy model of the crown’s lower ring, the “proto-crown.” And, inspired by this structure, the researchers began zeroing in on a short, rigid sequence of just 17 of protein A’s 998 amino acids that, like an arm crooked at the elbow, cradles other domains of the protein. The results are now out in the February 27 issue of Nucleic Acids Research.
“Despite its tiny size, the elbow became a surprising treasure trove of results, beautifully illustrating how combining structural results with other molecular biology approaches enables powerful studies,” says Ahlquist, the director of the John W. and Jeanne M. Rowe Center for Research in Virology at Morgridge.

Helena Jaramillo Mesa, a postdoctoral fellow in the Ahlquist Lab, led the investigation of the elbow region. The team suspected its position — at a crucial junction between subunits — may make it important to proto-crown assembly. What they unearthed were not just structural contributions but major responsibilities for turning on protein A’s two enzymatic functions: polymerase, essential for RNA synthesis, and RNA capping, essential for stability and expression of the new RNAs.
The study is the most detailed in vivo analysis of function in a viral RNA replication organelle to date. When they set out to mutate different segments of the 17 amino acids without disrupting the elbow structure, Jaramillo Mesa says, “we thought we were going to have some areas that were important, some that were not. But it turned out that when we mutated regions throughout the 17 amino acids, replication always went down.” The question became: “We know this region is important, but what exactly does it do?”
To answer this question, the researchers determined whether elbow mutants impaired RNA capping or polymerase activity or something else by testing whether they could “rescue” other mutants known to be defective in either capping or polymerase activity. Such rescue would suggest that an elbow mutant retained that particular function. “The big conclusion here is that specific elbow regions dramatically and selectively impact the enzymatic functions. And the most surprising thing is it doesn’t just impact the capping function that sits next to where the elbow is, but also the polymerase functions, which are much farther away from the elbow in the proto-crown conformation of protein A,” says Jaramillo Mesa.
Through further electron microscopy imaging, they tested whether mutations affected the assembly of the proto-crown and confirmed a third major role for the elbow: defects in the first 15 amino acids of the elbow led to microscopy images with no visible proto-crowns. “It is so surprising that different elbow regions have distinct, separable functions, or three different phenotypes, within just 17 amino acids,” Jaramillo Mesa explains. “It’s a master regulator and checkpoint for the whole system. Some of its interactions assemble the crown. Others link the completion of assembly to activating protein A’s RNA synthesis activities — ensuring that they act only in the right time and place for viral replication.”
Small elbow, big impact
While the nodavirus used in the study doesn’t pose a public health danger, it offers researchers an exemplary model system to discover fundamental principles. “Many insights from this work have implications for understanding and potentially controlling other viruses,” Ahlquist notes. Genes from this nodavirus also have advantages for potential beneficial applications of RNA replication, since “the gene encoding protein A is unusually small for its outsized role, and directs RNA replication in a remarkably wide range of cells.”
“It’s more important than ever to do this type of work because I think that all those public health decisions should ultimately be grounded in strong scientific evidence.” Helena Jaramillo Mesa
Indeed, the results of the study provide a clue into the evolution of the remarkably compact nodaviral genomes. Whereas many viruses expand their genome to encode proteases that cleave proteins so they can then take on different functions in replication, the nodavirus relies entirely on protein A, and in particular on the tiny but mighty elbow region.
Along with ongoing efforts to better understand a nearby flexible region adjacent to the elbow not yet visualized by cryo-EM, the study’s detailed insights into structure and function provide valuable foundations for opening new avenues in science and medicine. These include the development of broad-spectrum antivirals that target essential, conserved features shared across many viruses — features that are much harder for the virus to mutate under the genetic selection of antiviral treatment.
Jaramillo Mesa worries, however, about the challenge of communicating how such larger impacts emerge from fundamental research. “One thing we could do much better as scientists is communicating these ideas more clearly,” she says. “We’re working on very specific, very narrow things, so if we fail to explain how this impacts bigger questions, why would the public understand or trust us?”
This is a particular concern for virology, amidst shifting public opinion on vaccines and the research behind them. But Jaramillo Mesa is hopeful — and excited: “It’s more important than ever to do this type of work because I think that all those public health decisions should ultimately be grounded in strong scientific evidence.”
